FiberSim
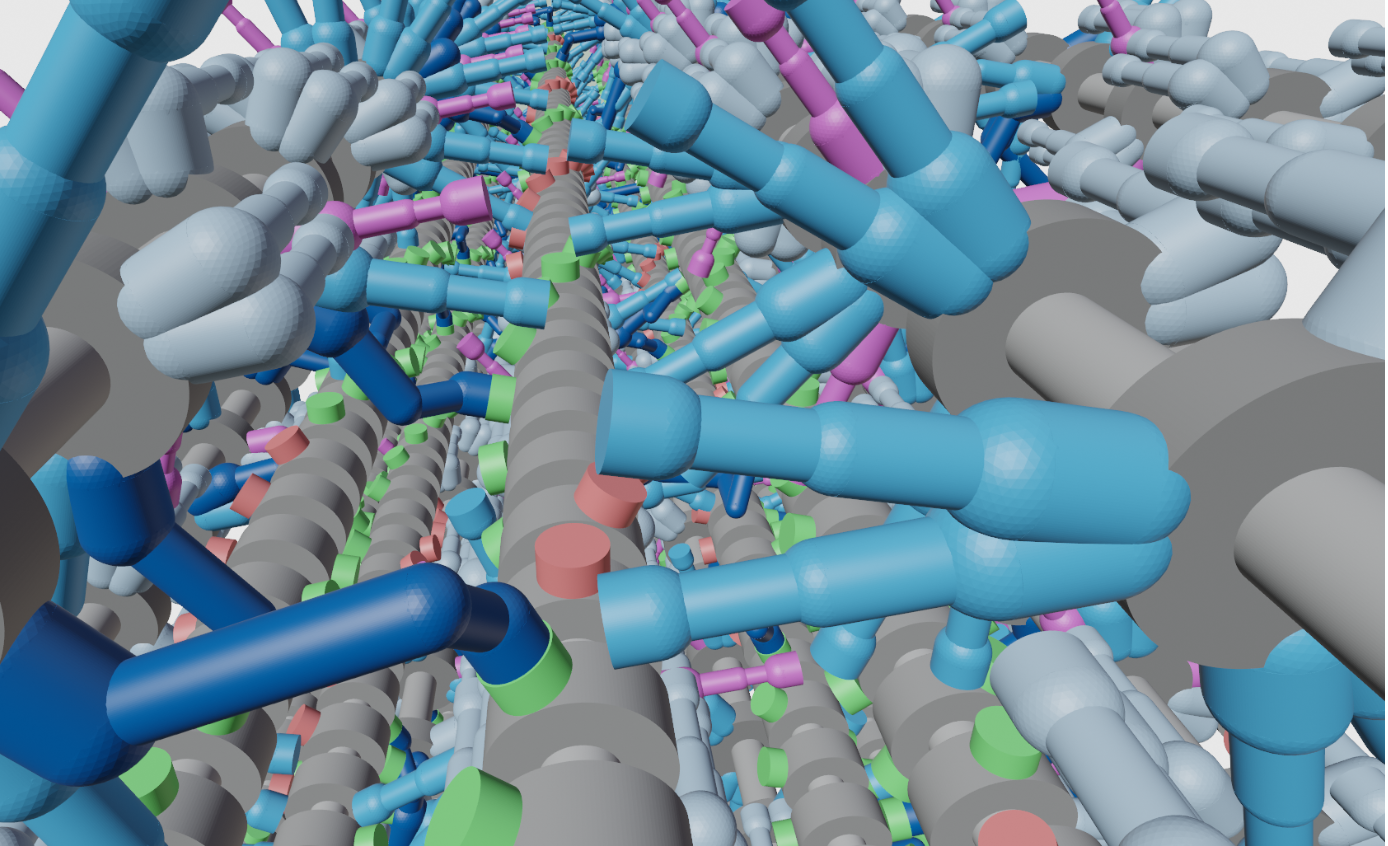
FiberSim is software for spatially-explicit modeling of half-sarcomeres. The code tracks the position and status of each myosin head, each binding site on actin, and each molecule of myosin binding protein-C.

You need a Windows PC to run a simulation but you can analyze output from the model on any computer that has an Anaconda-based installation of Python.
Organization
The main components of the software are:
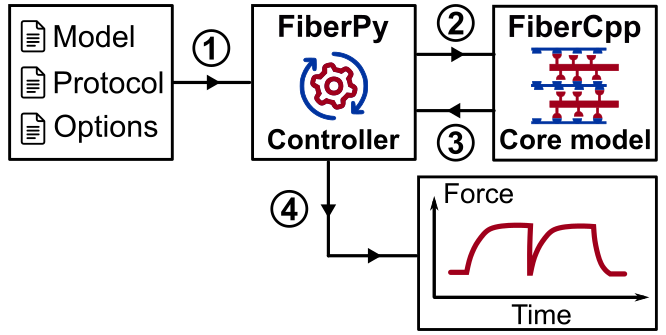
- FiberCpp - the core model that implements the calculations underlying the simulations.
This software is written in C++ but is currently only compiled for Windows PCs. - FiberPy - accessory code that makes it easier to run different types of simulations and analyze output.
This component is written in Python.
Getting started
Quick tutorial videos are provided here to help you install FiberSim. Then you can check the demos to see how to:
Run simulations Create videos and snap-shots of the model