Multiple half-sarcomere wtih series compliance
Overview
This demo shows how to simulate multiple half-sarcomeres that are connected in series with a linear spring.
What this demo does
This demo:
- Runs a single simulation in which a myofibril composed of 10 half-srcomeres connected in series with a linear spring is activated and deactivated by step changes in the Ca2+ concentration
- Plots summaries of the simulation
Instructions
If you need help with these step, check the installation instructions.
- Open an Anaconda prompt
- Activate the FiberSim environment
- Change directory to
<FiberSim_repo>/code/FiberPy/FiberPy - Run the command
python FiberPy.py characterize "../../../demo_files/myofibrils/multiple_hs_with_sec/base/setup.json" - You should see text appearing in the terminal window, showing that the simulations are running. When it finishes (this may take a few minutes), you should see something similar to the image below.
Viewing the results
All of the results from the simulation are written to files in <FiberSim_repo>/demo_files/myofibrils/mulitple_hs_with_sec/sim_data/sim_output
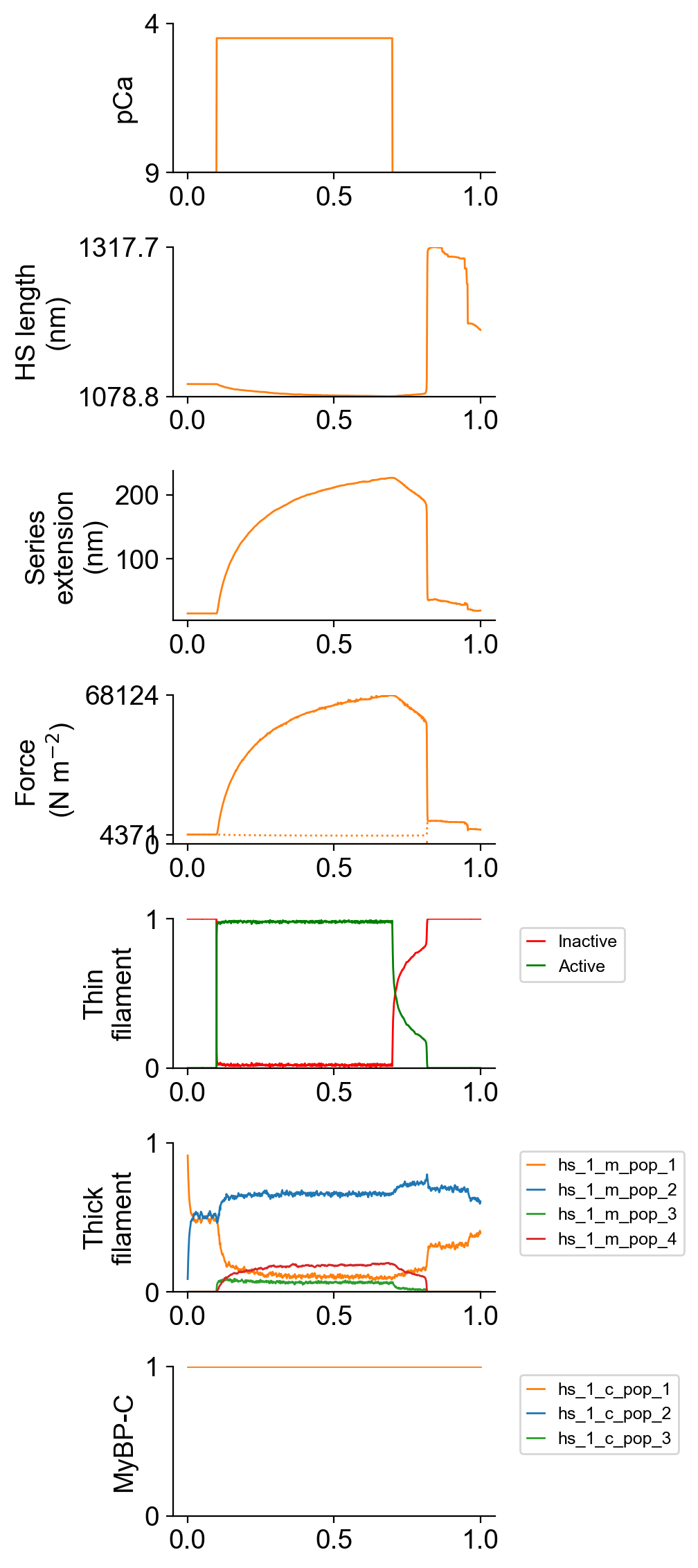
The file superposed_traces.png shows pCa, length, force per cross-sectional area (stress), and thick and thin filament properties for the first half-sarcomere in the myofibril plotted against time. Note the complex time-course of relaxation.
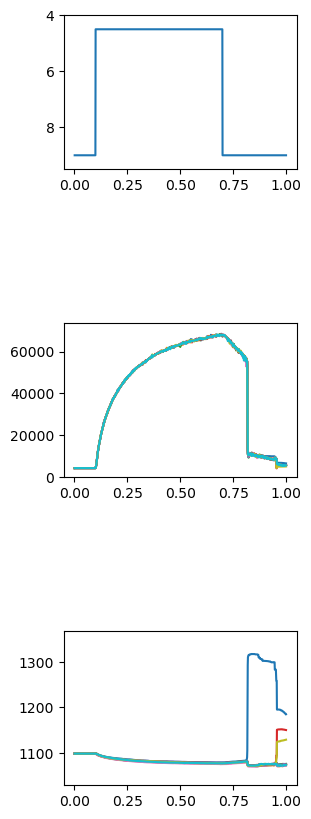
FiberPy also generated a customized figure for this simulation showing the force and length of each of the 10 half-sarcomeres in the myofibril. This file is named summary.png and is saved to the normal sim_output folder.
How this worked
The myofibril was defined by adding a series elastic stiffness sc_k_stiff to the muscle section and changing no_of_half_sarcomeres to 10 in <FiberSim_repo>/demo_files/myofibrils/hs_with_sec/base/model.json.
"muscle": {
"no_of_half_sarcomeres": 10,
"no_of_myofibrils": 1,
"sc_k_stiff": 3000,
"initial_hs_length": 1100,
"prop_fibrosis": 0.0,
"prop_myofilaments": 0.5,
"m_filament_density": 0.407e15
}
The characterization was the same as for the single half-sarcomere simulation except that a post_sim_Python_call was added. This points to a a standard Python file that is called after the simulation has finished. In this case, the Python made the summary.png figure showing the responses of the different half-sarcomeres.
{
"FiberSim_setup":
{
"FiberCpp_exe": {
"relative_to": "this_file",
"exe_file": "../../../../bin/FiberCpp.exe"
},
"model": {
"relative_to": "this_file",
"options_file": "sim_options.json",
"model_files": ["model.json"]
},
"characterization": [
{
"type": "pCa_length_control",
"relative_to": "this_file",
"sim_folder": "../sim_data",
"m_n": 9,
"pCa_values": [4.5],
"sim_duration_s": 1.0,
"time_step_s": 0.001,
"pCa_step_up_s": 0.1,
"pCa_step_down_s": 0.7,
"output_image_formats": [ "png" ],
"figures_only": "False",
"trace_figures_on": "False",
"post_sim_Python_call": "../Python_code/multiple_hs_summary.py"
}
]
}
}